|
For many RiPP families, a RiPP leader binding domain of the PqqD superfamily (PF05402) is found either as a discrete protein 9 or as a domain within tailoring enzymes, now referred to as a RiPP precursor peptide recognition element (RRE) 10. In response, numerous structures of RiPP modifying enzymes in complex with their precursors have begun to shed light on the contacts and strategies employed by RiPP biosynthetic enzymes. PTMs post-translational modifications, SAM S-adenosylmethionine, SAH S-adenosylhomocysteine. LC-MS/MS data show methylation of SonA-L63 always precedes methylation of SonA-I65, revealing N-to-C directionality for α- N-methylation as seen with all borosin methyltransferases to date (see also Supplementary Fig. d SonM-mediated α- N-methylation states of SonA. Deprotonation of the target amide hydrogen is proposed to be mediated either by an activated water 30 or conserved tyrosine 31. c General reaction mechanism for borosin precursor α- N-methylation. Type IV borosin pathways (reported in this manuscript) have distinct protein architectures, where dimeric α- N-methyltransferases associate with two discrete precursor peptides (no connective dashed segments). Types I–III borosin pathways 24 encode α- N-methyltransferases that are fused (dashed segments) to their precursor peptides, wrapping almost completely around the other subunit to achieve iterative intermolecular α- N-methylation. b Dimeric protein architectures of borosin α- N-methyltransferases. a General scheme for RiPP biosynthesis highlighting the typical composition of precursor peptides. Reaction schemes and protein structure configurations for the biosynthesis of borosin RiPP metabolites. The the combination of modifying enzyme promiscuity and regiospecificity has driven the field to seek structural clarity of these protein complexes and their underlying kinetic constraints to understand the rules governing RiPP PTM incorporation. In prokaryotes, RiPPs are often encoded in monocistronic, streamlined biosynthetic gene clusters where encoded tailoring enzymes frequently introduce multiple modifications in a given RiPP precursor 1. The short N-terminal leader peptides of RiPP precursors serve as docking domains that direct an assortment of tailoring enzymes to install post-translational modifications (PTMs) on the C-terminal core peptide destined to become the mature natural product (Fig. Initially translated by the ribosome, RiPPs are typically synthesized as short precursor peptides that are extensively posttranslationally modified at their C-termini prior to proteolytic maturation and export 1. 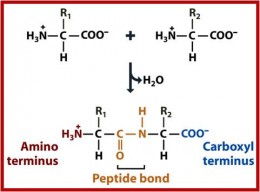
Moreover, peptide features such as amide backbone α- N-methylations 2, 3 and D-configured residues 4, 5, 6 once thought exclusive to non-ribosomal peptides are now known to also be installed by RiPP biosynthetic pathways. Over the past 25 years, ribosomally synthesized and posttranslationally modified peptides (RiPPs) have proven to be a major class of natural products, whose expanding breadth of unique structures and bioactivities are beginning to rival those of non-ribosomal peptides 1.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
